scRNA-seq Network Inference Benchmarking
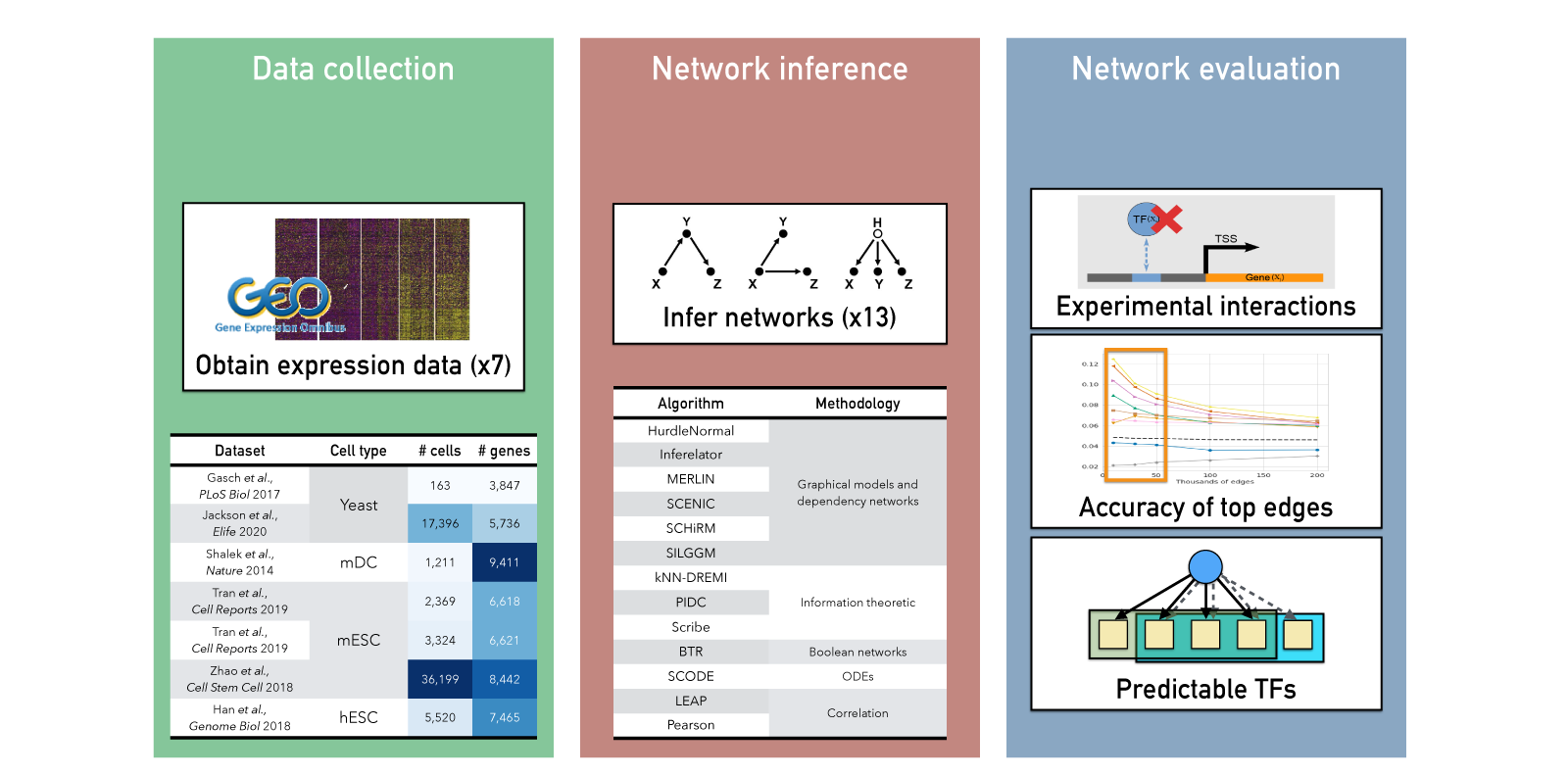
This project benchmarks gene regulatory network inference methods for scRNA-seq, comparing 11 algorithms across 7 published datasets (human, mouse, yeast) and multiple gold standards and metrics, including scalability (time, memory) and network recovery. Key findings: although overall recovery of global metrics (F-score and AUPR) is limited, methods capture biologically meaningful regulator–target interactions, and incorporating prior knowledge plus transcription factor activity estimation improves overall performance, while imputation did not help and can be detrimental.